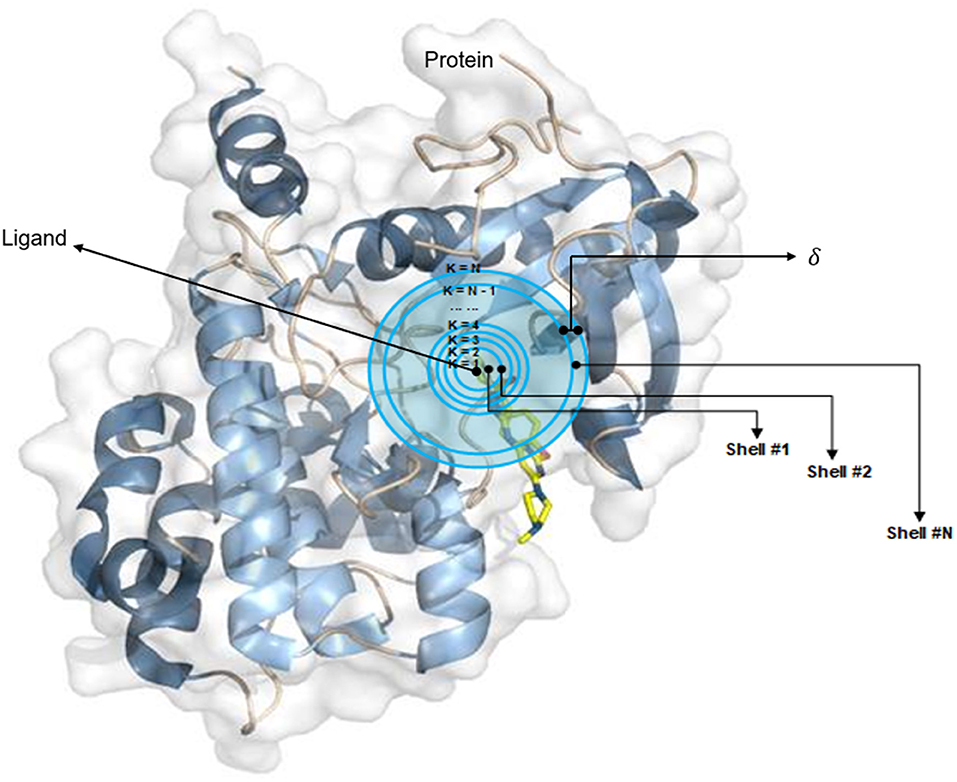
Frontiers | SE-OnionNet: A Convolution Neural Network for Protein–Ligand Binding Affinity Prediction | Genetics
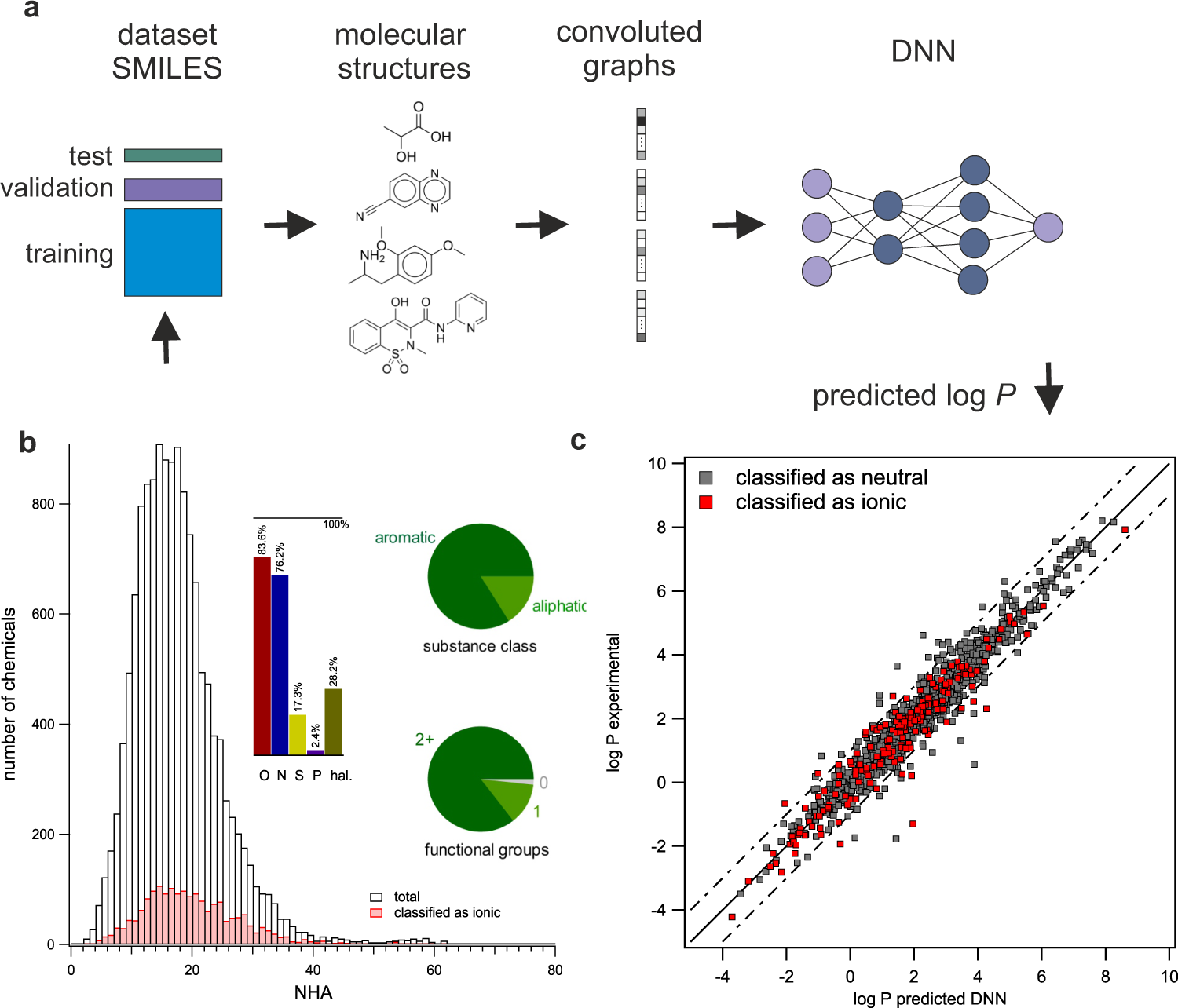
Exploring the octanol–water partition coefficient dataset using deep learning techniques and data augmentation | Communications Chemistry

Velocity prediction of nanofluid in a heated porous pipe: DEFIS learning of CFD results | Scientific Reports

Effect of Various Isolated Microbial Consortiums on the Biodegradation Process of Precipitated Asphaltenes from Crude Oil | ACS Omega

PDF) Efficient Toxicity Prediction via Simple Features Using Shallow Neural Networks and Decision Trees
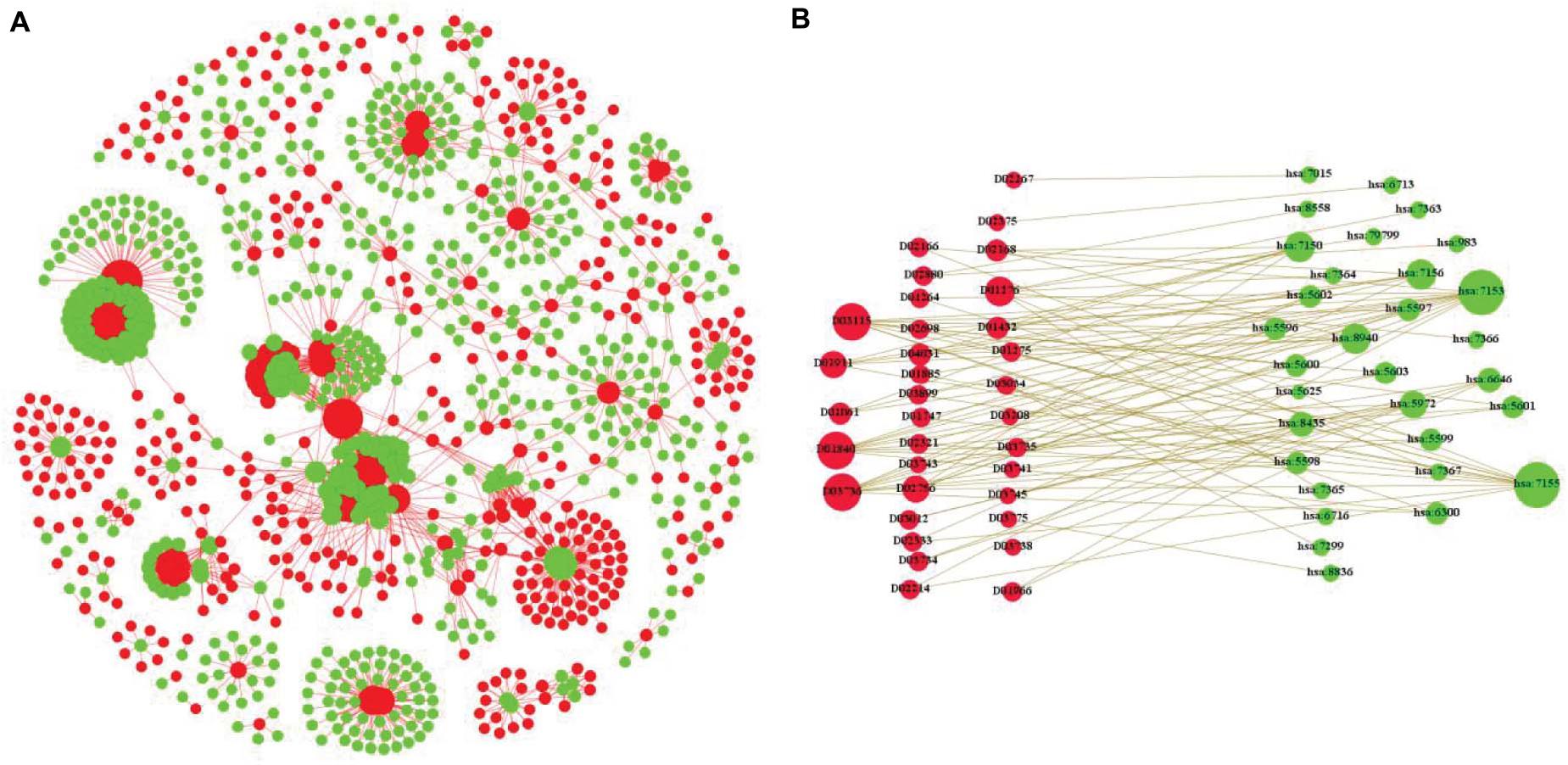
Frontiers | Cascade Deep Forest With Heterogeneous Similarity Measures for Drug–Target Interaction Prediction | Genetics
PDF) Dissolution Kinetics of a BCS Class II Active Pharmaceutical Ingredient: Diffusion-Based Model Validation and Prediction

Collaborative Approach between Explainable Artificial Intelligence and Simplified Chemical Interactions to Explore Active Ligands for Cyclin-Dependent Kinase 2 | ACS Omega
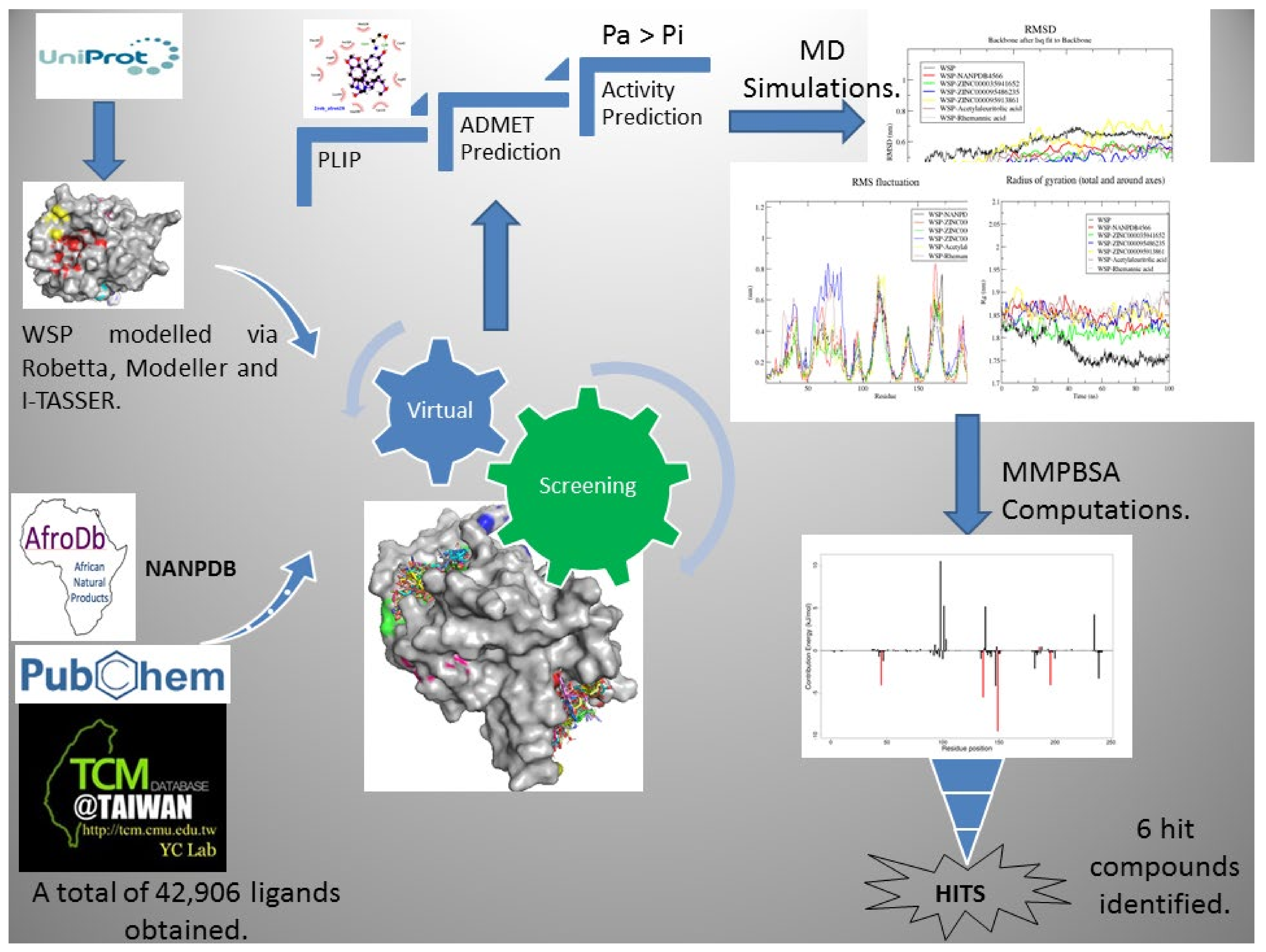
Biomedicines | Free Full-Text | Molecular Docking Simulation Studies Identifies Potential Natural Product Derived-Antiwolbachial Compounds as Filaricides against Onchocerciasis | HTML
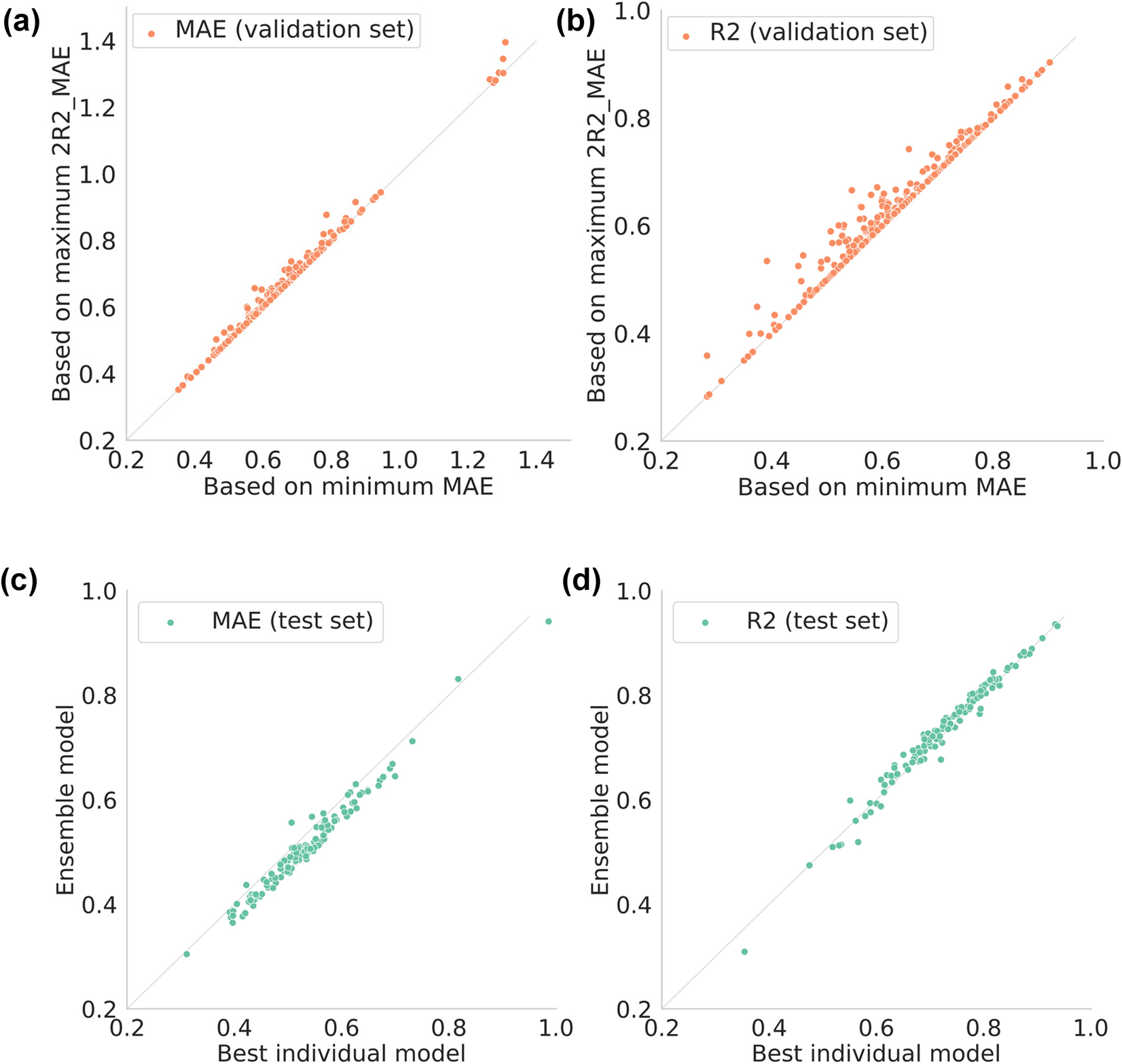
Prediction of pharmacological activities from chemical structures with graph convolutional neural networks | Scientific Reports
Machine learning-based solubility prediction and methodology evaluation of active pharmaceutical ingredients in industrial crystallization | SpringerLink

Machine Learning Approach to Predict the Surface Charge Density of Monodispersed Particles in Gas–Solid Fluidized Beds | ACS Omega